
Part 1: Implementing K-means clustering
In Part 1, you will design and implement a k-means clustering algorithm. You only need to modify kmeans.py for Part 1. Below is an overview of the data structures data and centroids, which you will need to work with in order to implement the algorithm:
data: A list of data points, where each data point is a list of floats.
Example: data = [[0.34, -0.2, 1.13, 4.3], [5.1, -12.6, -7.0, 1.9], [-15.7, 0.06, -7.1, 11.2]] Here, data contains three data points, where each data point is four-dimensional. In this example, point 1 is [0.34, -0.2, 1.13, 4.3], point 2 is [5.1, -12.6, -7.0, 1.9] and point 3 is [-15.7, 0.06, -7.1, 11.2].
centroids: A dictionary of centroids, where the keys are strings (centroid names) and the values are lists (centroid locations).
Example: centroids = {"centroid1": [1.1, 0.2, -3.1, -0.4], "centroid2": [9.3, 6.1, -4.7, 0.18]}. Here, centroids contain two key-value pairs. You should NOT change the names of the centroids when you later update their locations.
Info
When you first run the tests (before you implement anything) you will see several assertion errors. As you work your way through the assignment, you’ll start to see less errors as the tests pass. Seeing assertion errors for parts that you haven’t implemented yet is normal and expected.
We have provided several test functions that you should use to check your implementation as you go through the assignment. To run the tests, open the terminal (Terminal | New Terminal) and paste in python kmeans.py. You will only be working in kmeans.py for Part 1. Here is a brief overview of the steps, which are described below:
- Implement
euclidean_distance(), which will calculate the distance between two points. - Implement
get_closest_centroid(), which allows you to find the closest centroid for a point. - Implement
update_assignment(), which sets the centroid assignment for a point. - Implement
mean_of_points(), which calculates the mean of a list of points. Also, implementupdate_centroids(), which will update all of the centroids to the new mean of the points assigned to them. - Run the entire algorithm!
- Check code quality
- Submit your work!
Calculating Euclidean distances
Implement euclidean_distance() using the euclidean distance formula to calculate the distance between two points:
def euclidean_distance(point1, point2):
"""Calculate the Euclidean distance between two data points.
Arguments:
point1: a non-empty list of floats representing a data point
point2: a non-empty list of floats representing a data point
Returns: the Euclidean distance between two data points
Example:
Code:
point1 = [1.1, 1, 1, 0.5]
point2 = [4, 3.14, 2, 1]
print(euclidean_distance(point1, point2))
Output:
3.7735394525564456
"""
You should see the following output once you have correctly implemented euclidean_distance():
test_euclidean_distance passed.
This is what your terminal output should look like. Notice the AssertionError following test_euclidean_distance passed.; this is expected, as it comes from the test for the function you will implement in the next step. There is an error because you have not implemented it yet.
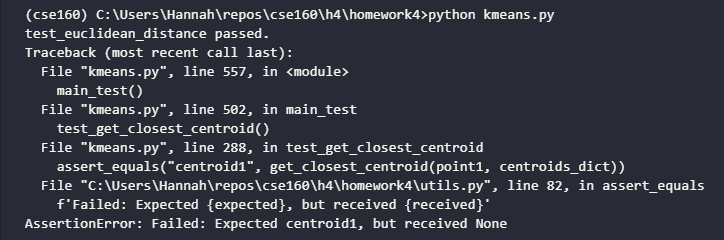
Assigning data points to closest centroids
Implement get_closest_centroid() to find the closest centroid for a data point:
def get_closest_centroid(point, centroids_dict):
"""Given a datapoint, finds the closest centroid. You should use
the euclidean_distance function (that you previously implemented).
Arguments:
point: a list of floats representing a data point
centroids_dict: a dictionary representing the centroids where the keys
are strings (centroid names) and the values are lists
of centroid locations
Returns: a string as the key name of the closest centroid to the data point
Example:
Code:
point = [0, 0, 0, 0]
centroids_dict = {"centroid1": [1, 1, 1, 1],
"centroid2": [2, 2, 2, 2]}
print(get_closest_centroid(point, centroids_dict))
Output:
centroid1
"""
You should see the following output once you have correctly implemented get_closest_centroid():
test_closest_centroid passed.
Again, you will also see error messages for the next tests, as you have not implemented those functions yet.
Update assignment of points to centroids
Implement update_assignment() to assign data points to their closest centroid:
def update_assignment(list_of_points, centroids_dict):
"""
Assign all data points to the closest centroids. You should use
the get_closest_centroid function (that you previously implemented).
Arguments:
list_of_points: a list of lists representing all data points
centroids_dict: a dictionary representing the centroids where the keys
are strings (centroid names) and the values are lists
of centroid locations
Returns: a new dictionary whose keys are the centroids' key names and
values are lists of points that belong to the centroid. If a
given centroid does not have any data points closest to it,
do not include the centroid in the returned dictionary.
Example:
Code:
list_of_points = [[1.1, 1, 1, 0.5], [4, 3.14, 2, 1], [0, 0, 0, 0]]
centroids_dict = {"centroid1": [1, 1, 1, 1],
"centroid2": [2, 2, 2, 2]}
print(update_assignment(list_of_points, centroids_dict))
Output:
{'centroid1': [[1.1, 1, 1, 0.5], [0, 0, 0, 0]],
'centroid2': [[4, 3.14, 2, 1]]}
"""
If there are no data points assigned to a centroid, then you should not include that centroid in the returned dictionary.
You should see the following output once you have correctly implemented update_assignment():
test_update_assignment passed.
Update centroids
Info
Do not modify list_of_points or hard-code the dimensionality of the data. Your code should be able to run on data points with any dimension.
Implement mean_of_points() to find the average of a cluster:
def mean_of_points(list_of_points):
"""Calculate the mean of a given group of data points. You should NOT
hard-code the dimensionality of the data points.
Arguments:
list_of_points: a list of lists representing a group of data points
Returns: a list of floats as the mean of the given data points
Example:
Code:
list_of_points = [[1.1, 1, 1, 0.5], [4, 3.14, 2, 1], [0, 0, 0, 0]]
print(mean_of_points(list_of_points))
Output:
[1.7, 1.3800000000000001, 1.0, 0.5]
"""
Then, implement the update_centroids() to update the centroid to be the average of the clusters:
def update_centroids(assignment_dict):
"""
Update centroid locations as the mean of all data points that belong
to the cluster. You should use the mean_of_points function (that you
previously implemented).
Arguments:
assignment_dict: the dictionary returned by update_assignment function
Returns: A new dictionary representing the updated centroids
"""
You should see the following output once you have correctly implemented mean_of_points() and update_centroids():
mean_of_points passed.
update_centroids passed.
At this point, if you completed all parts to the k-means clustering algorithm you should see following output when you run the tests:
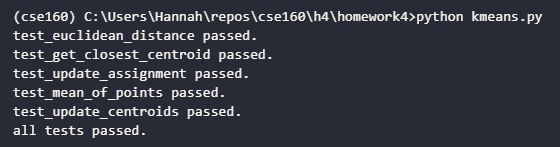
Clustering 2D points
Now that you have completed all parts needed to run your k-means clustering algorithm, we can now run it on actual data. At the very bottom of kmeans.py you should see code like this:
if __name__ == "__main__":
main_test()
# data, label = read_data("data/data_2d.csv")
# init_c = load_centroids("data/2d_init_centroids.csv")
# final_c = main_2d(data, init_c)
# write_centroids_tofile("2d_final_centroids.csv", final_c)
Comment out main_test() (which ran your tests) and uncomment the last four lines. These four will load the 2D data points and the initial centroids we provided in the .csv files. After that, main_2d() will execute the k-means algorithm and write_centroids_tofile() will save the final centroids to a file to save them permanently.
if __name__ == '__main__':
# main_test()
data, label = read_data("data/data_2d.csv")
init_c = load_centroids("data/2d_init_centroids.csv")
final_c = main_2d(data, init_c)
write_centroids_tofile("2d_final_centroids.csv", final_c)
Now run your program again by typing python kmeans.py in the terminal window. If you are successful, you should see:
K-means converged after 7 steps.
In addition, there should be 7 plots generated, in the results/2D/ folder. If you examine these 7 plots, they should be identical to the steps shown in the the gif shown in the background section of this spec.